
Why DNA Biomass ≠ True Abundance
DNA biomass is often mistaken for a direct measure of abundance, but eDNA levels fluctuate due to shedding, degradation, sampling and PCR bias. Here we explore why eDNA cannot quantify true abundance yet still provides robust, scalable insights for biodiversity monitoring.
By filtering water or air, collecting soil, or swabbing surfaces, we can detect hundreds of species without ever seeing a single organism. These developments mean eDNA methods have been enthusiastically adopted by ecologists because eDNA is so much better (in the current biodiversity crisis) at detecting low abundance species than many of the existing methods.
A common misconception is that the amount of DNA detected directly reflects the number of organisms present. It feels intuitive: more animals = more DNA. But that leap is scientifically weak. DNA biomass and true abundance can be linked, but imperfectly, inconsistently, and sometimes not at all.
The latest modelling framework for DNA-based surveys makes this clear:
“species-specific absolute abundances cannot be estimated using DNA data alone" (Diana et al., 2025)
Standard field surveys are not perfect they introduce bias through detectability, observer error, weather, activity patterns, and habitat accessibility. In a similar fashion, stages of the eDNA workflow can also introduce bias.
Organisms are known to shed DNA at very different rates. A European eel and a water vole, for example, contribute vastly different quantities of DNA to their environment. What's more within a population individuals will each contribute different amounts of eDNA dependant on physiology, size, stress, temperature, and behaviour. This variability means two populations with the same abundance can generate very different DNA signals.
Like a foggy day that impedes observations, environmental conditions can influence how long DNA persists. DNA degradation rates differ between habitats, seasons, temperatures, and substrates. A strong DNA signal can reflect presence, but it might also represent lingering DNA from an animal that is no longer there, or the rapidly degrading DNA from a species that is present in abundance. Results from one species or site therefore cannot be assumed to hold true elsewhere.
Observer bias is well documented in line transect surveys; the same bias can also be introduced with variable eDNA sampling techniques. Filtering 200 ml of river water versus 2000 ml, or swabbing tree roots rather than branches, will change both the species detected and the relative eDNA concentration.
DNA is composed of nucleotide sequences that are unique to each species, and these sequences amplify at different efficiencies. Some species’ DNA binds better to primers, outcompetes others in PCR, or amplifies earlier. The result is PCR bias, where a species that was relatively rare in the environment may appear abundant in sequencing data, or vice versa. This is why metabarcoding read counts cannot be treated as direct abundance estimates.
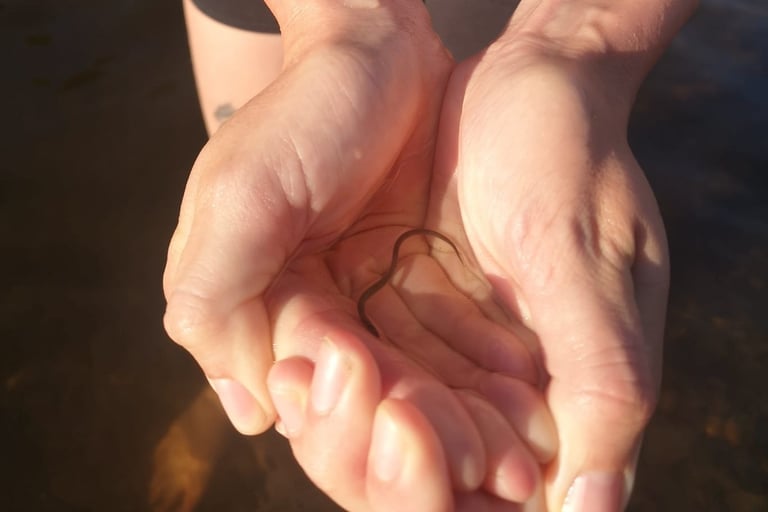
With rigorously standardised field and laboratory workflows, it is sometimes possible to infer relative changes in DNA biomass for a single species across sites (Diana et al., 2025). This works best with targeted, single-species assays, where no other species’ DNA is competing within the PCR. Even then, results must be interpreted with caution and ideally calibrated against independent ecological data. Weldon et., al (2020) developed an eDNA assay for European eel that is now widely used across the industry. The assay was validated in Irish lochs working with the eel monitoring team at Inland Fisheries Ireland. The study found single species eDNA and qPCR could show the relative abundance of eel populations between sites in a single survey season.


It is worth highlighting that in most real-world ecological monitoring, true abundance is rarely measured. As a result, many abundance estimates end up binned into categories like high, medium, and low, or converted into indices that serve as proxies rather than literal counts. In practice, conservationists are almost always asking bigger questions than “How many individuals exist?” They want to know whether a population is healthy, whether a site is functioning normally, or whether a protected area is delivering conservation outcomes.
The goal, in other words, is to assess population and ecosystem health, not to count every individual.
This is exactly where eDNA is already proving its value. Even without absolute abundance, eDNA can provide robust indicators of community change, species turnover, and early warning signals. With the rapid emergence of new tools such as environmental RNA (eRNA) and using eDNA detections in combination with functional groups, we are now moving beyond simple presence/absence towards detecting activity, life stage, and improvements in functional processes.
So while eDNA cannot deliver true abundance, the ecological world rarely operates on that level anyway. When used correctly and interpreted cautiously, eDNA can answer the same core questions ecologists have always cared about, and often more efficiently and at greater scale than traditional methods ever could.
Alex Diana, Eleni Matechou, Jim Griffin, Douglas W. Yu, Mingjie Luo, Marie Tosa, Alex Bush & Richard A. Griffiths (2025) eDNAPlus: A Unifying Modeling Framework for DNA-based Biodiversity Monitoring, Journal of the American Statistical Association, 120:549, 120-134, DOI: 10.1080/01621459.2024.2412362
Weldon, L., O’Leary, C., Steer, M., Newton, L., Macdonald, H. and Sargeant, S.L., 2020. A comparison of European eel Anguilla anguilla eDNA concentrations to fyke net catches in five Irish lakes. Environmental DNA, 2(4), pp.587-600.
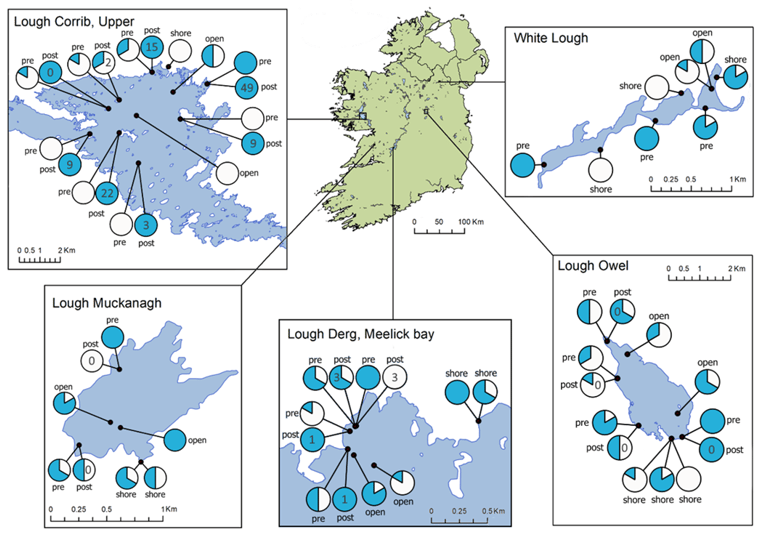
Anguilla anguilla detections from Weldon et al., (2020)
Anguilla anguilla.
The eDNA Consultancy
Where our expertise meets yours
ADDRESS
Unit 1, Tanhouse Farm, Frampton on Severn, Gloucestershire, GL2 7EH, GB
INQUIRE ABOUT OUR SERVICES
Web design and graphics by Bear Bones
The eDNA Consultancy Ltd is registered in England and Wales 15508607. VAT no: 482885735 © Copyright The eDNA Consultancy Ltd. All rights reserved.
